The following is adapted from an August 2025 response I wrote to a creationist on X who goes by the handle “@DivinelyDesined“, henceforth “DD”. I put a fair amount of work into it and don’t think it got a sufficient hearing then/there, so I repost it here with some editing and additions for a hopefully wider audience. My apologies if some of this gets a bit technical, I had to read up a bit on this subject myself to formulate my response, so wherever technical terms pop up I have tried to link to relevant explanatory resources. Also, DD’s writing is a little scattered and repetitious. I tried to organize the primary issues into the same piles, but it is sometimes difficult, hopefully things aren’t too hard to follow.
DD, sadly, has over 12k followers on X and frequently posts “long” (for X) and very self-assured, dismissive attacks on evolutionary theory, largely made up of misinformation and strawman caricatures of what evolution. For example, DD once complained about an illustration showing an example of macroevolution, in this case a species of butterfly giving rise to a new species of butterfly, by suggesting that a better example of macroevolution would be “…a butterfly transforming into a dragonfly.” Anyone with an even basic understanding of evolutionary theory knows that it does not postulate that species from one existing clade will somehow morph into species of a completely different clade (see monophyly). This is not how evolution works; this is not what the basis of macroevolution (speciation) looks like. And in the case of butterflies and dragonflies, neither group is descended from the other, rather they share a common ancestor that while it was an insect was neither a butterfly nor a dragonfly.
In another DD posts with apparent approval a page out of an antievolution cartoon tract by the late Jack Chick titled “Bid Daddy” (pp.12-13) This alone is enough to discredit DD as a serious person, but never mind.
This particular engagement started with one of DD’s responses to an evolution defender on X who goes by the handle “Creationist Translationist” [@JustinCPorter], henceforth “CT”:
DD: The argument is not that there is a mechanism which stops genetic variation or change from accruing — the argument is that the change which accrues cannot ever build novel biological structures necessary for macroevolutionary innovation. [Link to source, 06/20/2025]
After which DD gave the following link to another thread where DD had stated the following:
DD: There is no evidence that random changes in DNA will ever add up over time to construct novel genes & cellular structures. [Link to source, 03/07/2025] …The evolutionist believes that mutations will add up over time and engineer novel genes, which leads to novel proteins, cells, organs, and body plans – despite the fact that we have ZERO evidence of this ever occurring, nor even being possible. [Link to source, 03/07/2025]
CT responded with a reiteration of one of his earlier responses:
CT: Without a mechanism to stop variation from accruing iver [sic, “every” -TB] generations, you have nothing to prevent speciation from continuing over millions — even billions — of generations. You are a fake skeptic. [C.T. on X 06/20/2025]
And I jumped in and added to the thread an admittedly lazy, throw away response, of the following:
Me: That and: [Google Scholar link to papers on de novo genes coding for proteins.]
DD responded to me with the following:
DD: Let’s see if you understand what you’re reading, and not just forming opinions based on misunderstanding hyped up headlines. Pick a paper from that list – any paper – and summarize why it provides evidence for Evolutionism. [From DD’s response to my throw away.]
To which I replied:
Me: LOL! This is not my first rodeo. I am not going to fall for the beach bum creationist routine where I do a bunch of work presenting evidence that the claims you make are nonsense, only to have you, based on willful ignorance and an existential dread of actually understanding the thing you’re sure isn’t so (evolution), dismiss whatever I show you as insufficient. For example, you have in the past said that macroevolution would be something like a “butterfly transforming into a dragonfly”, something which is impossible according to evolutionary theory. So, I could show you legit evidence of macroevolution, as the term is defined by scientists, and you would simply reject it out of hand based on it not matching your twisted creationist definition of the term. How about this instead? You’re claiming that the vast literature on the genetics of evolution is fundamentally flawed, and that the overwhelming majority of geneticists don’t know what they’re talking about, how about YOU pick a (recent, say from the last 10 years) paper and walk us through where they are factually incorrect or logically incoherent.
“Evolutionism.” 🙄 [Link to source, 06/21/25]
DD responded not by addressing any of the papers I had linked from Google Scholar, or even any similar paper dealing with the subjects DD laid out in his earlier comments about “novel genes & cellular structures” or “novel proteins, cells, organs, and body plans” but rather with a repost of something DD had written earlier attacking a paper on an RNA replication experiment, which is related to prebiotic evolution (abiogenesis/RNA world hypotheses) rather than biological evolution proper which was, as I understood it, what DD’s bluster regarding “novel genes” etc. was about.
I guess it’s partly my fault for not being more specific about the subject matter of the paper DD could pick. I thought it should have been obvious from the context, but oh well.
A quick side note for DD here; when you are going to critique a paper or article like this, you really should provide a link so that people can go and look at the material for themselves without having to manually type in the title from a screen shot into a search engine. It is common curtesy for your readers and fair play for the target of your criticism.
That is unless your intent is to discourage your readers from looking at the material and have no interest in fair play. I mean this paper isn’t even behind a paywall. Here are a text-based title and link to the paper that will be under discussion:
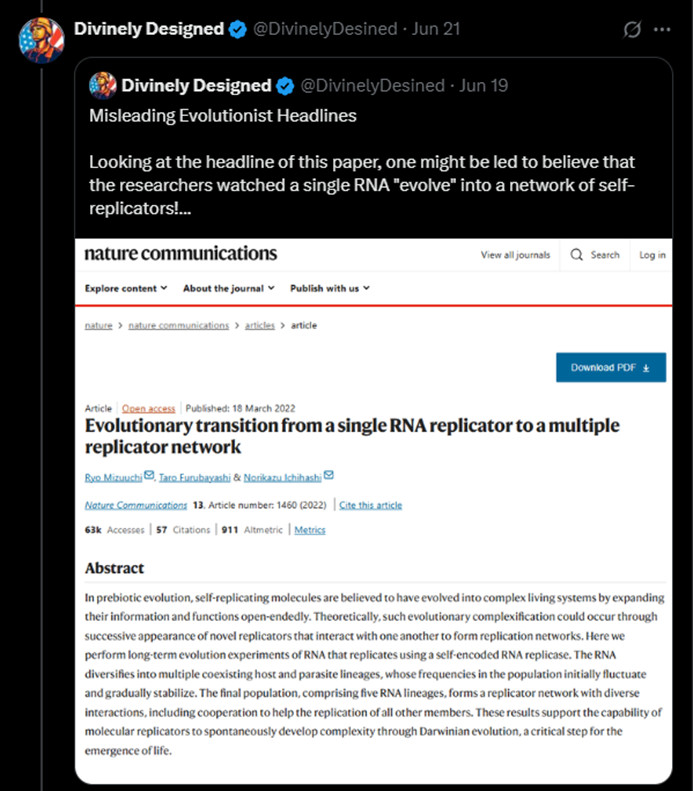
Mizuuchi, R., Furubayashi, T. & Ichihashi, N. (2022) “Evolutionary transition from a single RNA replicator to a multiple replicator network” in Nature Communications 13:1460 doi.org/10.1038/s41467
And additionally here is a link to supplementary material the authors also published:
So now, even though this is on a different subject—prebiotic “evolution” rather than biotic evolution (evolution proper)—we are going to take a look at DD supposedly showing us where these authors were “factually incorrect or logically incoherent” (which is what I suggested DD do) and some of his responses to my initial questions regarding said “analysis”.
DD: Misleading Evolutionist Headlines Looking at the headline of this paper, one might be led to believe that the researchers watched a single RNA “evolve” into a network of self-replicators!
DD: In this experiment, researchers set out to investigate if a single RNA strand could evolve into a network of cooperating molecules.
In a way, they did accomplish this, but the results are not nearly so exciting for Evolutionism as many Evos believe. [Link to source, 06/19/2025]
They accomplished what they were trying to do, so how is the title, “Evolutionary transition from a single RNA replicator to a multiple replicator network” misleading then? Is it misleading just because the results, in your oh so objective “judgment”, weren’t as exciting as some supposed unnamed evolutionists supposedly believe?
Who are these supposed evolutionists that are more excited about this paper than they should be? Who DD? Give us some names. And what is it that they are too excited about? You’ve already admitted that the researchers accomplished what they were trying to do.
Or is this just rhetorical nonsense?
Personally, though I thought it was interesting, I didn’t get all excited about it, but then I am not a molecular guy.
DD: The researchers in this experiment started out with a single RNA strand, which was specifically designed & engineered to copy itself. [Link to source, 06/19/2025,Emphasis in original.]
DD: This RNA could never exist in nature – it can only survive under strict lab conditions. [Link to source, 06/19/2025,Emphasis in original.]
DD: The researchers also used an RNA Replication System – also specifically designed & engineered to replicate this RNA strand. [Link to source, 06/19/2025, Emphasis in original.]
DD: The RNA strand was able to catalyze the RNA Replication System, kickstarting the replication process. [Link to source, 06/19/25]
The RNA replication system (Fig. 1a) consists of a single-stranded RNA (host RNA) that encodes the catalytic subunit of Qβ replicase (an RNA-dependent RNA polymerase) and a reconstituted Escherichia coli translation system. RNA replication occurs through the translation of the replicase subunit, which becomes active in association with elongation factors Tu and Ts (EF-Tu and EF-Ts) in the translation system. [Mizuuchi et al (2022) p.2 (emphasis mine)]
DD: So, first thing to ask is, was this a “self” replicating RNA?
I would argue no. The RNA catalyzed its replication in another replication system.
So it didn’t actually replicate itself – it simply turned on the Copy Machine. [Link to source, 06/19/2025]
Again, the RNA did not act as a catalyst in this case. It encoded a protein, Qβ replicase, that acted as a catalyst. Ribosomes “borrowed” from E. coli found in the “translation system” read the original RNA which encoded the Qβ replicase protein and produced them. Once these proteins were made, they started replicating the RNA strands leading, ultimately, to the five primary RNA lineages (3 host & 2 parasites).
DD: In the process of replication, there were mutations.
These mutations led to shorter RNA strands than the original – notice how the mutations were DELETERIOUS in nature, not adding anything new, but instead degrading the original information. [Link to source, 06/19/2025]
DD: Some of these new, shorter strands took on specialized roles within a cooperative network of other strands. Instead of one Copy Machine, now it had many parts working together to do the same task. [Link to source, 06/19/2025]
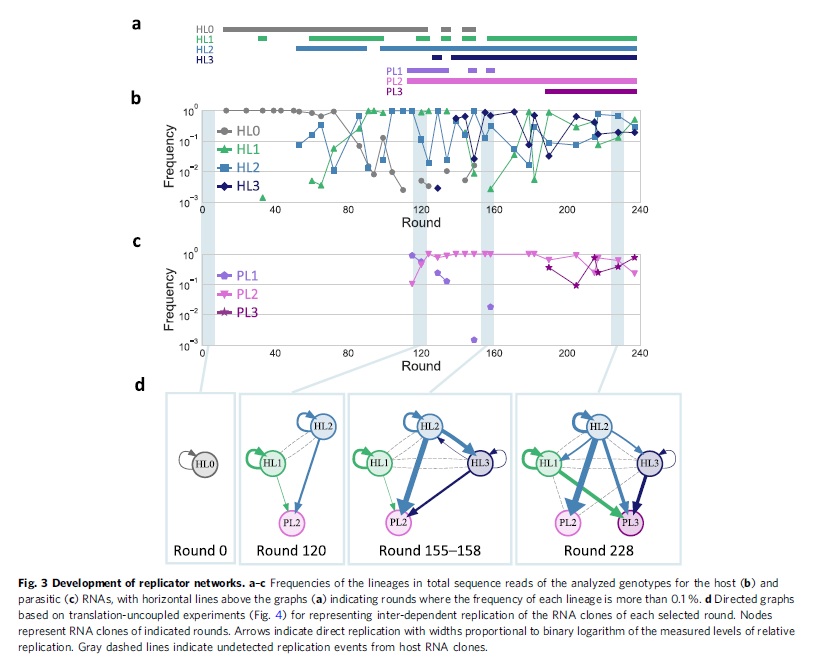
The mechanism that sustained the replication of multiple RNAs is not obvious from the structure of the replicator network (Fig. 3d) because it lacks direct reciprocal interactions between replicators, which have been employed in both simulations and experiments to demonstrate the coexistence of multiple molecular replicators. [Mizuuchi et al (2022), p.7 (emphasis mine)]
So, no “reciprocal interactions” between the replicators; they did not cooperate in that sense. The presence or absence of individual lineages did affect the other lineages, but not due to some sort of cooperation toward some common goal. Also, the only “copy machines” were the ribosomes in the translation system (to read the host RNAs and create Qβ replicase) and the Qβ replicase that copied the RNA.
Next, DD essentially accuses the authors of the paper of putting their thumbs on the scale of the experiment by interfering with it in such a way as to achieve a desired result:
DD: HOWEVER – some of these mutated strands acted like “parasites” and caused problems in the system by interfering with the replication process.
These problematic RNAs either competed for resources or disrupted the cooperative system, destabilizing it. [Link to source, 06/19/2025]
The experimental setup required that certain steps be taken to prevent parasitic lineages from prematurely crashing the ecosystems (more on this shortly), all of which is explained in the article, so nothing underhanded. However, at the end there were 3 hosts and 2 parasitic lineages which were all part of a relatively stable ecosystem (“network”):
Our results demonstrate an evolutionary transition scenario of molecular replicators from a single common ancestor to a multi-membered network. During long-term replication, a clonal host RNA population gradually diversified into multiple host and parasitic RNA lineages. Their frequencies initially fluctuated probably due to competition, but stabilized as the evolution progressed (Fig. 3a–c). The co-replication of isolated RNAs in five lineages exhibited the gradual development of replicator networks, eventually consisting of five types of RNAs with distinct features (Fig. 3d). [Mizuuchi et al (2022), pp. 6-7]
DD continues:
DD: To maintain the experiment’s focus on the cooperative network, the researchers actively removed or suppressed these troublesome RNA strands.
Read that again.
The researchers REMOVED any PROBLEMS from their experiment. [Link to source, 06/19/2025]
Me: Where does it say that they “removed or suppressed” the parasites to “maintain the experiments focus”? Which is it btw, remove or suppress? How did they suppress them? Did they remove all of them? [Link to source, 06/22/2025]
DD: On Page 2, they state, “When encapsulated in micro-sized water-in-oil droplets, host RNAs can sustainably replicate in the presence of parasitic RNAs.”[Source: DD’s response to my clarification questions. 06/25/2025]
What the authors are referring to in what DD quotes here is part of the initial setup of the experiment. It involves creating a mixture of water and oil. The original RNAs and translation system were placed in water and then mixed with oil such that you end up with little bubbles of oil encapsulating droplets of water containing the RNA mixture (this roughly simulates lipid encapsulation of a hypothetical protocell).
This was done because when you put the RNA mixture in one container of water, the parasites tend to quickly evolve and collapse the entire RNA ecosystem. By breaking up the mixture into numerous water droplets encapsulated by oil, they created numerous tiny ecosystems. This allowed for some mini ecosystems to develop where the parasites, by chance, had not collapsed the system, because the host RNAs in those mini ecosystems adapted to their parasites (by making variations of the Qβ replicase that either do not replicate parasite RNAs or at least do so less efficiently than host RNAs). They then, as shown in Figure 1, took a sample out of the mixture, diluted it with fresh translation system, and placed it in a new container of oil and “…vigorously mixed to induce their random fusion and division.”
If you want to characterize that as the authors “suppressing” the parasites, sure I guess, but what they did is stated explicitly, and justified in the paper and isn’t some some sort of cheating that DD implies with his dramatic ALL CAPS.
DD: This encapsulation compartmentalized parasitic RNAs, allowing the host RNAs to persist. Page 7: “One key factor may be a compartment that could prevent the complete domination of parasites…”[Source: DD’s response to my clarification questions. 06/25/2025]
It encapsulates all the RNAs—the original RNA strand only initially, and then different mixtures of host and parasite RNAs—into numerous water-in-oil droplets. It does not sequester parasite RNAs completely from the hosts. Again, it merely creates lots of mini ecosystems rather than one big one.
None of this involves the active removal of parasites from the experiment that DD accused the authors of doing. It only prevents them from dominating the early stages of the experiment until the hosts can evolve resistance in some of the encapsulated droplets.
Now back to the subject of the earlier all caps claiming that the authors of the paper “REMOVED any PROBLEMS from their experiment.”
DD: They also removed replicating RNAs from the central pool to observe them independently of the system at different times during the experiment. [Source: DD’s response to my clarification questions. 06/25/2025]
NO THEY DIDN’T (see I can all caps to).
They did not “remove” the parasites. DD’s own quote from the paper earlier in response to my question states that the hosts were still “in the presence of the parasites”.
Further, if one reads the paper carefully, they will see that the only actual “removal” of different RNA strands was done in a computer simulation run parallel with the physical experiment! Quote:
Next, to understand whether the replication relationships between these RNA clones (Fig. 3d) could sufficiently explain the observed replication dynamics, we created a theoretical model of the five RNAs whose replication rates were determined from average replication levels of HL1-, HL2-, HL3-, PL2-, and PL3–228 in the translation-uncoupled RNA replication experiments (Fig. 4). We simulated continuous replication in uniform-sized compartments by modeling the three steps shown in Fig. 1b. Consistent with the experimental results, four RNAs, except for one based on HL3–228, sustainably co-replicated by displaying similar concentration dynamics (Fig. 5b). These experimental and simulation results demonstrated that the replication relationship among the selected RNA clones explain the sustainable replication of at least four RNA lineages.
Finally, we examined how the four RNAs could sustainably replicate when competing with each other. Hypothesizing that each RNA helped sustain the replication of others, we simulated continuous replication by removing each of the four RNAs from the five-member network. We found that the absence of HL1-, HL2-, PL2-, and PL3–228 tended to cause the extinction of PL3-, PL2-, HL1-, HL2-, and PL2-228, respectively (Supplementary Fig. S21). A serial transfer experiment in the absence of HL2–228 reproduced the simulated replication dynamics in that PL2-228 was diluted out (Supplementary Fig. S20b). These results support the interdependence of the four RNAs, facilitating the coexistence of multiple replicators. [Mizuuchi et al (2022) pp. 5–6 (emphasis mine)]
Of note, the experiments in the absence of the other RNAs (HL1-, PL2-, and PL3–228) were unsuccessful because the removed RNAs appeared soon, probably due to the presence of contamination at an undetected level and/or de novo production through recombination and mutations. [Mizuuchi et al (2022) p.6]
The mechanism that sustained the replication of multiple RNAs is not obvious from the structure of the replicator network (Fig. 3d) because it lacks direct reciprocal interactions between replicators, which have been employed in both simulations and experiments to demonstrate the coexistence of multiple molecular replicators12–16,31. One key factor may be a compartment that could prevent the complete domination of parasites in the population 28,32,33. Moreover, our simulation displayed that removing one RNA replicator typically drove other RNAs to extinction (Supplementary Fig. S21), partly supported by experiments (Supplementary Fig. S20b), indicating that the RNAs in the network may have sustained each other indirectly. The mechanisms behind these observations could be explained as follows. (1) Removal of HL1–228 resulted in the disappearance of PL3–228 through competition with PL2-228, which adapted to HL2–228. (2) Removal of HL2–228 caused the extinction of PL2-228 because it could not replicate in the absence of HL2–228. (3) Removal of PL2-228 led to the competitive exclusion of HL1–228 by HL2–228, which replicated slightly faster and was more resistant to PL3–228. (4) Removal of PL3–228 made HL1–228 outcompete HL2–228 because the remaining parasitic RNA (PL2-228) parasitized only HL2–228 among the host RNAs. The disappearance of HL2–228 then caused the extinction of PL2-228, as its replication relied on HL2–228. Overall, all RNAs aided the replication balance of each other, and thus, the long-term coexistence. [Mizuuchi et al (2022), p.7 (emphasis mine)]
Finally on the issue of selective lineage removal, I took the trouble to contact one of the authors (Norikazu Ichihashi, personal communication 06/2025), who confirmed that they never intentionally removed any RNAs from the system in the main experiment and that the sequential removal of different types of RNA discussed in the paper was only done in the simulation. So, DD’s implied accusation regarding the authors’ supposed fixing of the experiment by removing parasitic lineages to get some predetermined result also seems to be baseless.
Now back to the RNA strands supposedly all getting shorter.
DD: They started out with one RNA Replicase, and over time, it “evolved” into an RNA Replicase made up of many smaller RNA strands instead of one large one. [Link to source, 06/19/2025]
No, they started out with on type of RNA that encoded Qβ replicase, which allowed the initial lineage of one type of RNA to evolve into multiple lineages of RNA, some shorter (parasites) and others roughly maintaining the original length (hosts).
In the paper under discussion the authors do talk about three lineages of RNA that acquired mutations deleting their genes for Qβ replicase causing them to become parasitic on the remaining “host” lineages (only two of these parasitic lineages, PL2 & PL3, survived to the end of the experiment); “parasitic” in this case means the parasite RNAs were reliant on the Qβ replicase still coded for in the host RNAs. However, DD doesn’t make any such distinctions. His is a blanket claim that the RNAs got shorter compared to the original RNA placed in the experiment.
So, once again I asked DD for clarification on this:
Me: Where in the paper does it say that the descendant host RNA strands got shorter? [Link to source, 06/22/2025]
DD: On Page 2, the researchers state explicitly that, “Spontaneous RNA recombination deletes a part of the replicase gene to generate parasitic RNAs while retaining 5′ and 3′ terminal sequences for recognition by the replicase.” [Source: DD’s response to my clarification questions. 06/25/2025]
DD: On Page 3 they mention that the “Sequence analysis uncovered that two previously decreased host RNA lineages became sustained…” [Source: DD’s response to my clarification questions. 06/25/2025]
Here, we continued the serial transfer experiment up to 240 rounds (1200 h). Sequence analysis uncovered that two previously detected host RNA lineages became sustained and further diverged into multiple sublineages of host and parasitic RNAs. The population dynamics of each lineage gradually changed during the evolution, from dynamically fluctuating stages to quasi-stable coexistence, suggesting the appearance of co-replicative relationships among the lineages. [Mizuuchi et al (2022), p.2 (emphasis mine)]
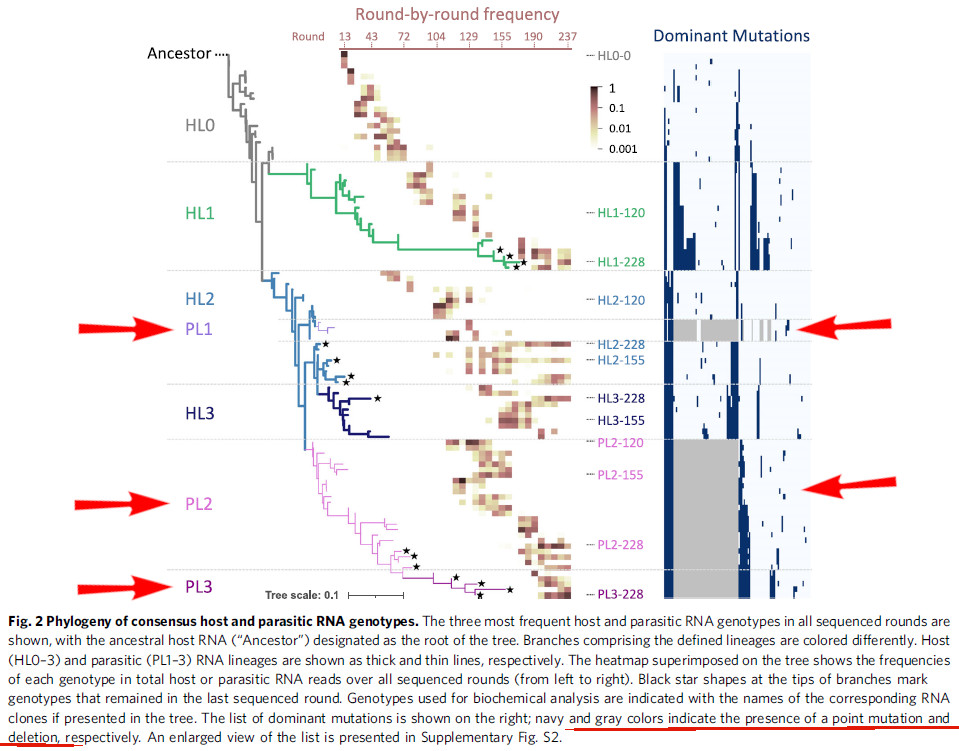
DD: The ensuing discussion implies that lineages contain shorter variants. The phylogenetic tree (Fig 2) and mutation list (Supplementary Fig S2) detail specific deletions, with different RNAs shown to have accumulated deletions, reducing their length compared to the ancestral host RNA. [Source: DD’s response to my clarification questions. 06/25/2025]
There is nothing that I can find in the paper stating that the host lineages in the experiment got significantly shorter due to deletions. I ran the paper through both ChatGPT and Grok AIs; both confirmed my reading about the host lineages. And once again I took the trouble to contact one of the authors of the paper (ibid.), and he confirmed for me that the size of the host lineage RNAs remained almost constant and that there was no trend for them to become shortened.
So, I have no idea where DD is getting the idea that the RNAs were “degrading,” unless DD could not tell the difference in the paper between the host and parasite lineages of RNA. There could also be some reading into the text of the paper what DD wants to see, i.e., confirmation of the belief that all mutations lead to degradation of some sort.
DD: So, over time, instead of a single RNA doing all the work, these shorter, helpful variants that were not artificially removed, evolved to form a system where they helped replicate each other. [Link to source, 06/19/2025]
DD: The researchers didn’t actually see anything new evolve.
They started out with one RNA Replicase, and over time, it “evolved” into an RNA Replicase made up of many smaller RNA strands instead of one large one. [Link to source, 06/19/2025]
DD: And it was ONLY able to get as far as it did because the researchers continuously removed anything that slowed it down.
As we have already see, this is false.
DD: I would argue this paper absolutely contradicts Evolutionism, as stated previously. Instead of evolving functional innovation, we get parasites and deletions.[Source: DD’s response to my clarification questions. 06/25/2025]
Evolutionary theory (“evolutionism” isn’t a thing) deals with the change and diversification of life on Earth through time. This is an experiment dealing with replicating RNA molecules, not living things; they’re not even RNA viruses. There appears to have been some type of selection processes going on in interactions of the RNA lineages, that I would suggest is only tangentially related to biological evolution proper. It has more bearing on abiogenesis research, a separate field of research from evolution.
And even with that, DD mischaracterizes the results of the experiment.
The experiment started with a single host lineage of RNA which developed into multiple new lineages of both host RNAs and parasitic RNAs (the only ones with large-scale deletions). Then these new RNAs—hosts and parasites—formed an interdependent relationship where none had existed previously.
How does this contradict evolution again? Evolution does not predict that all changes will be additive or always increasing in complexity. As stated earlier, sometimes, especially with parasites, organisms greatly simplify because some of their former complexity is no longer needed to survive and reproduce.
That said, there has been a general trend toward greater complexity in life on Earth. Life during most of this planet’s history consisted of nothing more than single-celled bacteria/archaea → then eukaryotes evolved with their more complex structures → then multicellular forms evolved → then more complex plants and animals. This is not a necessary prediction of evolutionary theory; it is an observation from the fossil record.
DD: The end system still did the same thing as the original, but with many pieces instead of one whole strand, and that was only accomplished because of heavy investigator interference.[Source: DD’s response to my clarification questions. 06/25/2025]
What the system did throughout was make copies of RNA. The result wasn’t the same as the original, and there was never just one “whole” strand. At the start there was a population of the modified original host RNA (coded to produce Qβ replicase), which turned into three separate host lineages—about the same as the original in size—and two shortened parasitic lineages (which lost their Qβ replicase coding).
And once again, DD’s accusation of “heavy investigator interference” appears to be based entirely on DD’s misreading of the paper and a deep seated desire to see malfeasance on the part of scientists that isn’t there.
DD: If conditions were anything close to the natural world, this experiment would have showed complete destruction of the entire system within a few rounds.[Source: DD’s response to my clarification questions. 06/25/25]
DD: The only thing this study shows is intelligent design, genetic entropy, and deleterious mutations. [Link to source, 06/19/2025]
We then plotted the frequencies of each lineage for every sequenced round (Fig. 3). For the host RNA lineages (Fig. 3b), the ancestral lineage, HL0, gradually became less frequent from round 72. Instead, the frequencies of HL1, HL2, and HL3 increased but highly fluctuated ranging from the undetected level to almost 100% until round 190. Afterward, the frequencies of all three evolved lineages became relatively stable as they were persistently detected at more than 3% of the population. For the parasitic RNA lineages (Fig. 3c), PL1 was initially dominant but quickly became rare. In contrast, PL2 dominated the population throughout the rounds with more than 10% of the population. PL3 became dominant at round 190 and then coexisted with PL2 at similar frequencies. Overall, the five RNA lineages, HL1, HL2, HL3, PL2, and PL3, were initially rare or highly fluctuated in frequency but shifted to relatively stable coexistence in the last ~50 rounds of the experiment, as illustrated in Fig. 3a. [Mizuuchi et al (2022), p.3]
A) Yes, there was “intelligent design” in setting up the experiment: the scientists involved were examples of embodied human intelligence, the only relatively advanced technological intelligence currently known to exist. We have no evidence for the existence of a supernatural disembodied intelligence that exists outside of spacetime, and no way to test for such a thing’s existence.
B) “genetic entropy” is a creationist invention—particularly of retired Cornell University geneticist John C. Sanford—
which Sanford describes thusly:
What is Genetic Entropy? It is the genetic degeneration of living things. Genetic entropy is the systematic breakdown of the internal biological information systems that make life alive. Genetic entropy results from genetic mutations, which are typographical errors in the programming of life (life’s instruction manuals). Mutations systematically erode the information that encodes life’s many essential functions. Biological information consists of a large set of specifications, and random mutations systematically scramble these specifications — gradually but relentlessly destroying the programming instructions essential to life. — Sanford
And finally, C): if there were only deleterious mutations, how could the population of RNA strands in the experiment have gone from one parent strand to being replaced by multiple host and parasite lineages that together formed a stable ecosystem? If “genetic entropy,” with only “deleterious mutations,” were the only active mechanisms acting on the experimental population, why would separate lineages develop? And shouldn’t the lineages, once somehow developed, have simply petered out to extinction rather than developing a persistent, more complex (compared to the original) interactive system?
DD’s claims regarding “genetic entropy” and “deleterious mutations” simply do not follow from the data presented in the paper.
DD: Which is EXACTLY what the Creationist model predict we would find in life. [Link to source, 06/19/2025]
DD: Funny how Evolutionist experiments actually provide evidence for Creationism. [Link to source, 06/19/2025]
DD: Now, please make a respectful argument about why my interpretation is wrong.[Source: DD’s response to my clarification questions. 06/25/25]
Says the person whose every scribbling on X, about evolution and those who study it, drips with contempt.
Considering all this I don’t believe anyone should be taking Divinely Designed frequent and grandiose claims to having debunked evolution (or any science) seriously.
DD is All hat and no cattle. All bark and no bite. All foam and no beer. All show and no go. All sizzle and no steak.
You get the picture.
Note: As of the time of this posting Divinely Designed has not responded to the above comments originally posted on X, despite my tagging him in the opening post of the thread. DD claimed in one of the back-and-forth exchanges between us that they get a lot of messages and sometimes (conveniently?) misses’ things, so after posting this I will tag DD again on X and point to this posting to give him another opportunity to not “miss” it.